Viral diagnostics
Viral Diagnostics
Cube Dx offers a fast yet comprehensive mutational analysis of SARS-Cov-2. The fast analysis enables a swift stratification of patients and a fast reaction on new variants, popping up the first time in a region. A detailed mutation pattern furthermore facilitates the choice of samples that will be further analysed with NGS. Mutations especially in the receptor binding domain (RBD) of the S-gene are relevant as these changes might hamper the effectiveness of vaccines, the clinical presentation of the disease and diagnostic tools.
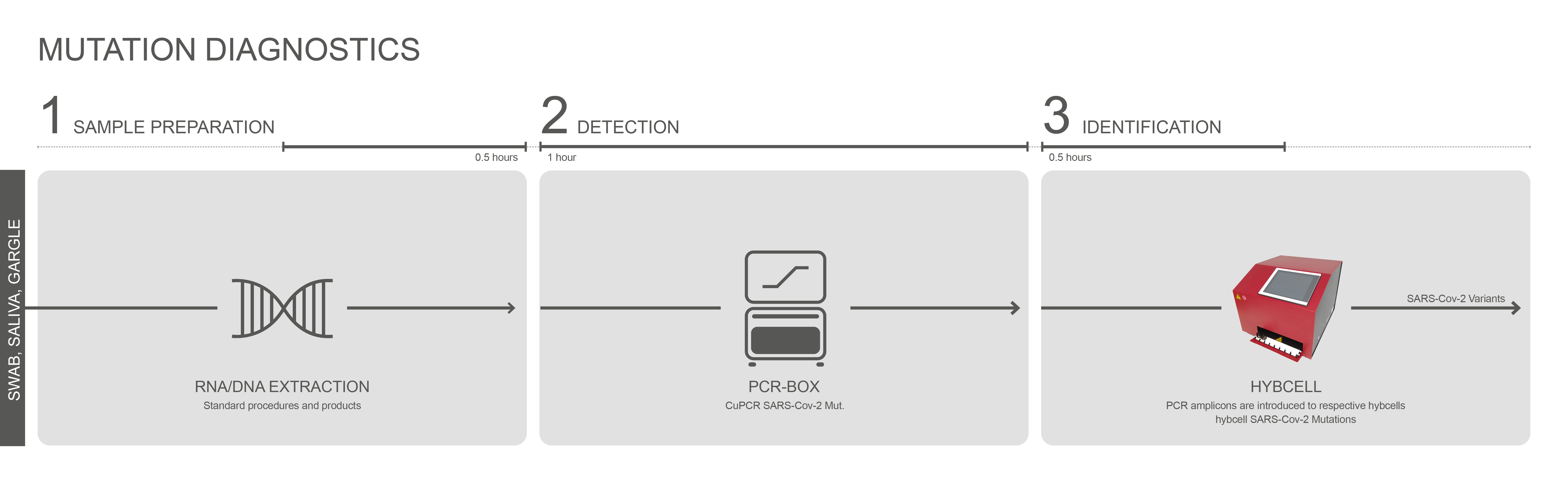
Compact sequencing is a two-step process. The first step is a standard RT- PCR reaction (of a fragment of the S-gene). This first step provides single-stranded fluorescence-labelled DNA for the second step and allows for a quantitative estimate of viral load. The second step is the identification of mutations and data analysis in the hyborg. Combining a quantitative estimate of viral load and at the same time testing for different and parallel variants (mixtures) is unique. The simple handling, the comprehensive mutation panel, and the automated analysis of variants drastically reduce the need for personnel for mutation diagnostics. The whole process takes less than two hours.
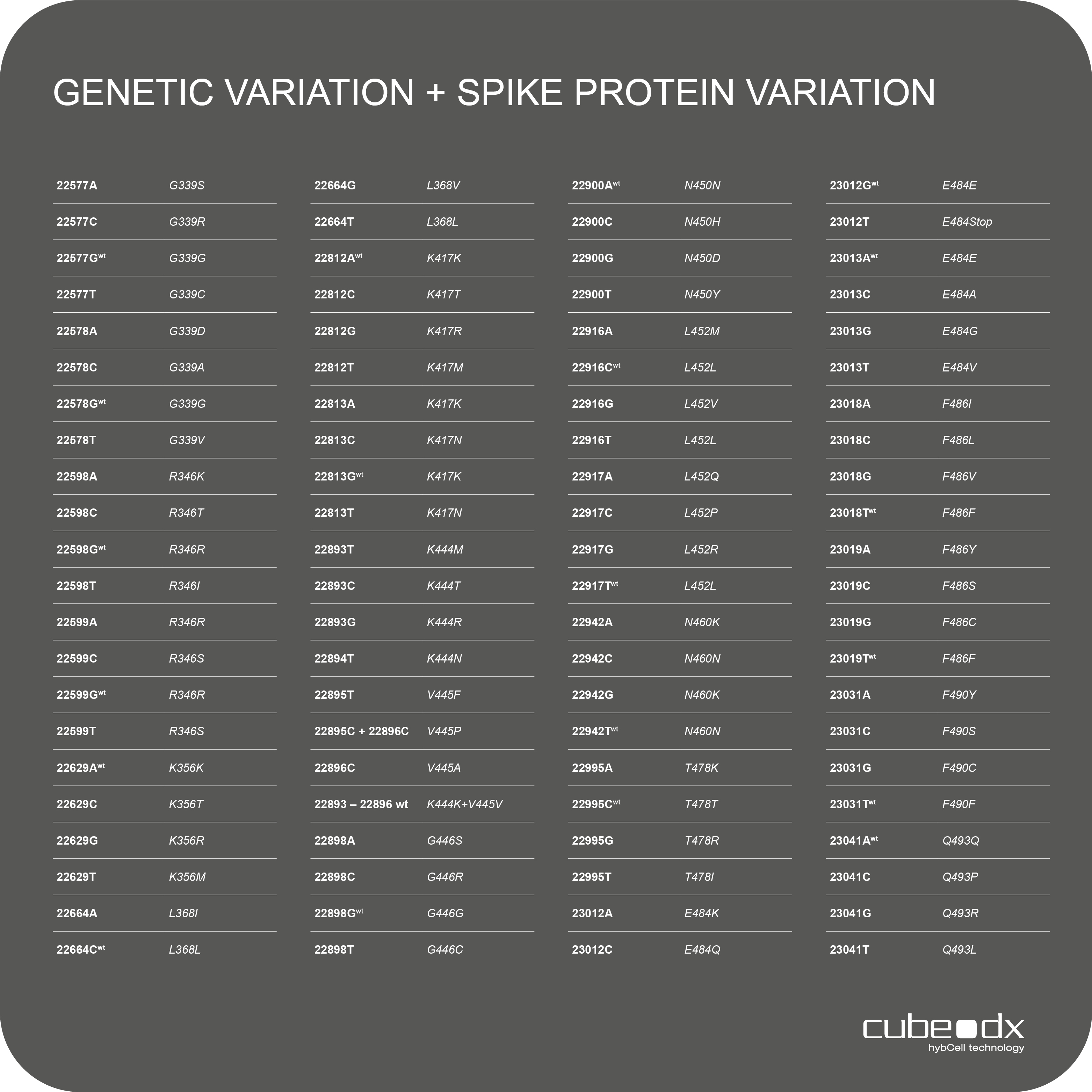
88 mutations / All relevant Omicron variants
EPIDEMIOLOGICAL SURVEILLANCE OF SARS-COV-2 MUTATIONS/VARIANTS:
Cube Dx’ fast and comprehensive examination of mutations of SARS-Cov-2 quickly singles out virus phenotypes with altered mutation patterns to take swift action to contain an outbreak of new variants.
The multiplex RT-PCR CuPCR SARS-Cov-2 Mutations is the first step for the mutation analysis and is based on a one-step reaction of reverse transcription (RT) combined with real-time PCR for the detection of the RNA of SARS-Cov-2 (S gene). The test is based on the principles of TaqMan-PCR and uses a single fluorescence dye to indicate amplification of the S-gene.
Positive amplicons are immediately examined for relevant mutations in the receptor binding domain (RBD) of the S gene of SARS-Cov-2 with hybcell SARS-Cov-2 Mutations using Cube Dx’s proprietary compact sequencing. In just 15 minutes, 88 SNPs (single nucleotide polymorphism = mutations) can be distinguished by identifying bases (A, C, G, T) of +ssRNA at specific nucleotide positions. The resulting mutation patterns are translated into likely variants of concern (or interest) by the software.
The test is periodically updated to cover the currently most relevant mutations and variants. Different samples can be tested, for example human samples or wastewater samples. The test is an epidemiological tool and is not intended to serve as an in-vitro diagnostic tool.
Advantages:
-
Screening of 1 amplicon (S-gene)
-
Minimum hands-on time, and direct use of positive amplicons, fast results, and "easy" read-out of results (report)
-
Limit of detection > 10 RNA copies per reaction
-
Analysis of 88 SARS-Cov-2 SNPs in 20 minutes
-
Automated reporting of known variants without any user intervention
-
Ability to show different variants simultaneously in aggregate samples (e.g. wastewater)
-
Validated for human and (waste) water samples